What
The MLCommons Medical Working Group continues to advance real-world benchmarking and evaluation of AI/ML models in healthcare with MedPerf, its open-source orchestrator designed for federated studies. Building upon expertise on global large clinical studies, MedPerf is now integrating Apache Airflow to make data preparation pipelines distribution, orchestration and monitoring in federated studies much easier. By combining MedPerf’s federated experiment orchestration with Airflow’s mature scheduling and monitoring capabilities, the Medical WG makes it easier for the community to run reproducible clinical AI studies in the real-world.
Why
The need for Airflow integration arose from challenges we faced when running our past clinical studies. Before this integration the Benchmark Committee on MedPerf was in charge of creating a single data preparation container, which was then shipped to the Data Provider to run data preparation locally. This monolithic containerization approach often led to unexpected errors, such as Data Providers having different types of input data than expected or Data Providers confusing execution instructions. Troubleshooting also proved challenging, as the pipeline developers would have to remotely assist users. Moreover, a single container for the entire data preparation process made it impossible for Data Providers to easily resume after an error. Finally single pipeline steps cannot be reusable in other pipelines. During the Federated Learning for Postoperative Segmentation of Treated glioblastoma (FL-PoST) study, a single container was developed that could resume execution after being interrupted (please see our previous technical report for details on this). While this approach solved most issues, it was not scalable because new containers would have to be built for each new study.
Solution
The Airflow integration on MedPerf aims to improve data preparation usability and robustness in a scalable way. Thanks to Airflow, it is possible to chain multiple containers into a single Data Preparation workflow. This brings two key benefits to MedPerf users:
- Each container becomes a modular step of the Data Preparation pipeline, allowing it to be utilized in multiple different workflows
- Airflow has its own WebUI for pipeline execution and monitoring, which makes it easier to diagnose errors and allows users to restart pipelines from arbitrary steps.
To make things much easier for MedPerf non-technical users (i.e. study coordinators), they can build their own pipelines using a single YAML file (examples are provided here). The MedPerf client automatically converts this into executable Airflow’s Directed Acyclic Graphs (DAGs) so that Airflow can execute it.
Technical Details
Whenever MedPerf instantiates Airflow it runs only locally, using a randomly generated password each time, and is shut down automatically once data preparation is complete or interrupted by the user. This ensures protection of sensitive data on the Data Provider’s side. The integration also includes a small utility that periodically reports data preparation status back to the MedPerf server using Airflow’s REST API. This is very beneficial to the Benchmark Committee to review the data preparation status across the whole federation (i.e. multiple sites) and provide proactive assistance to the participating sites.
Example: Data Preparation on Federated Learning for Postoperative Segmentation of Treated glioblastoma (FL-PoST)
The new implementation of the data preparation pipeline for Federated Learning for Postoperative Segmentation of Treated glioblastoma (FL-PoST) study is described in detail on the README.md file in the MedPerf code repository. Below we describe key points of this implementation.
The status for data preparation can be easily verified in the Airflow’s WebUI. Figure 1 shows the status of data preparation (i.e. step) on a single site through Airflow’s WebUI, while Figure 2 shows a partial graph view displaying the first few steps of the workflow.
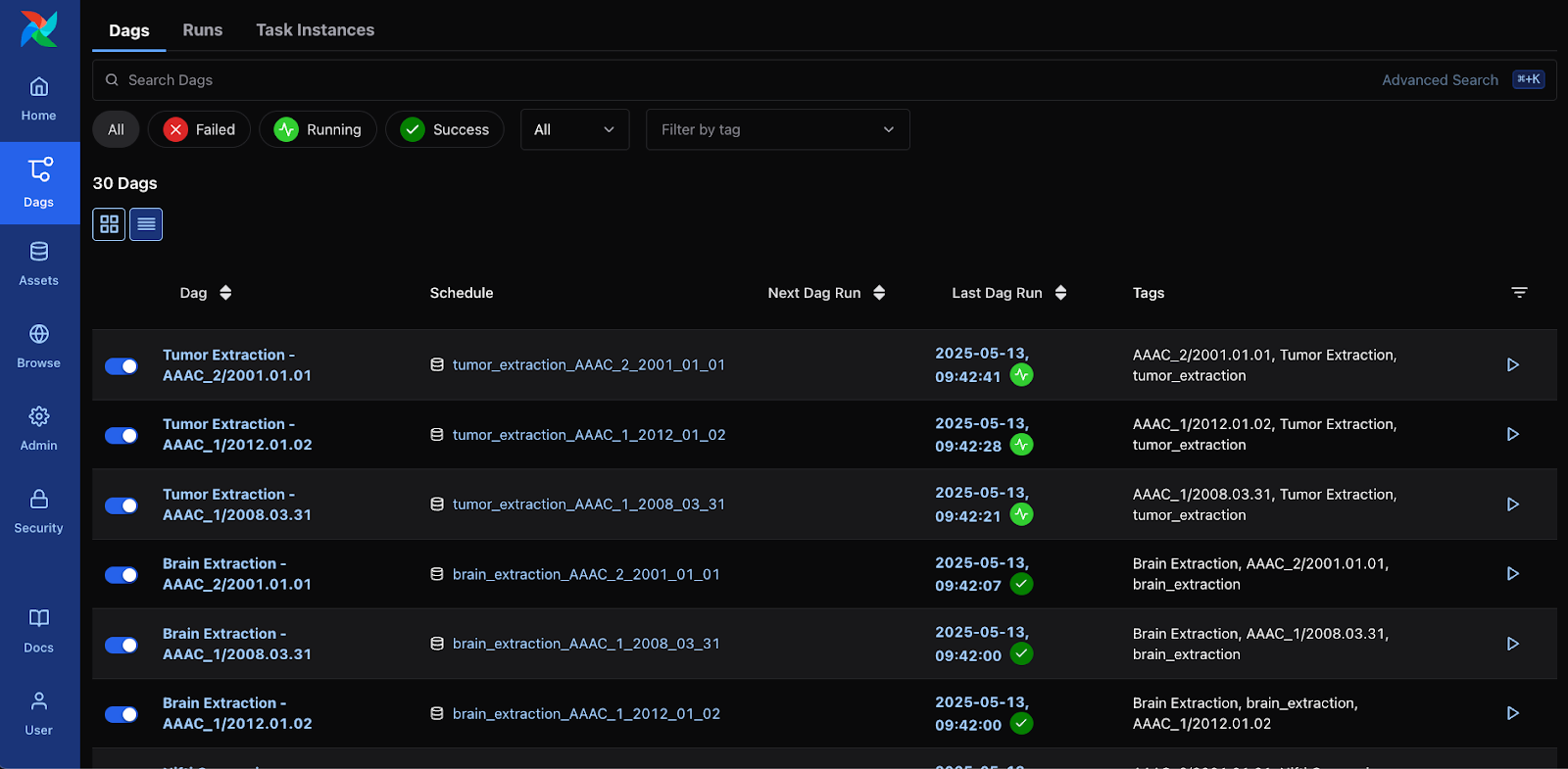
Figure 1: DAG view on Airflow, corresponding to each step of the data preparation along with its status
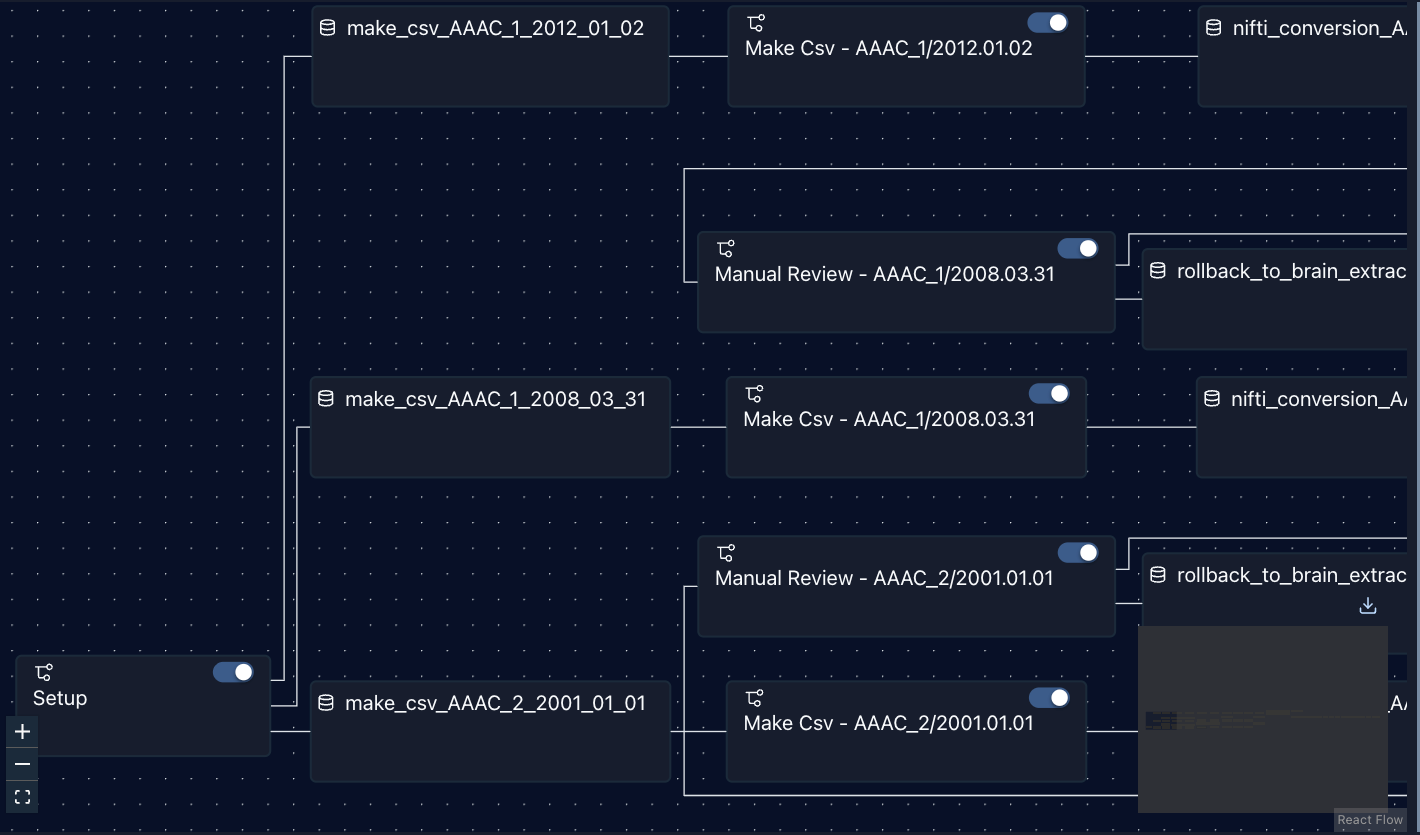
Figure 2: Partial Graph View of all generated tasks in Airflow.
The data preparation pipeline of this particular study also includes a Manual Review step, in which a human operator on the Data Provider’s side must manually review the generated tumor segmentation maps and approve or make further annotation corrections. Meanwhile Airflow sets the task into an “Up for Reschedule” and waits until the file is reviewed and copied into the proper directory per data preparation guidelines (see Figure 3). Once all reviewed files are reviewed and copied per data preparation guidelines, Airflow will automatically resume execution.
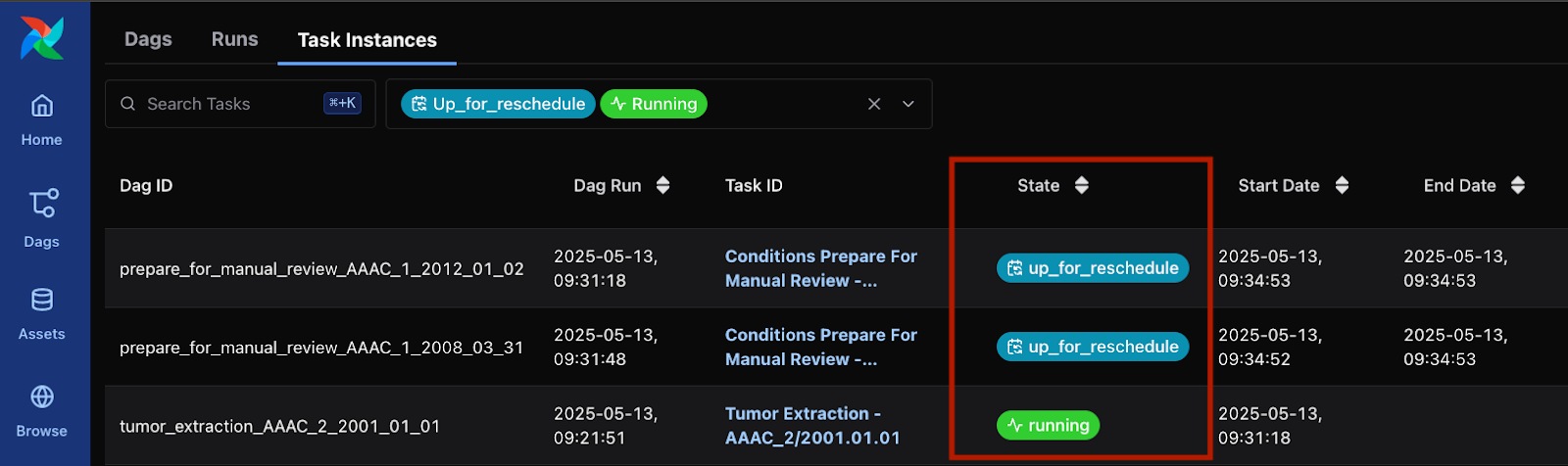
Figure 3: Airflow waiting for Manual Review in two tasks, while running a different task
Airflow’s “Approval Operator” feature, as of version 3.1.0, enables a human operator to issue a manual approval for pipelines to run. In the Federated Learning for Postoperative Segmentation of Treated glioblastoma (FL-PoST) study such a manual approval is requested once all brain scans have been processed and reviewed by the human operator (see Figure 4). This is the final step after which the pipeline automatically generates the train and test splits for the experiment.
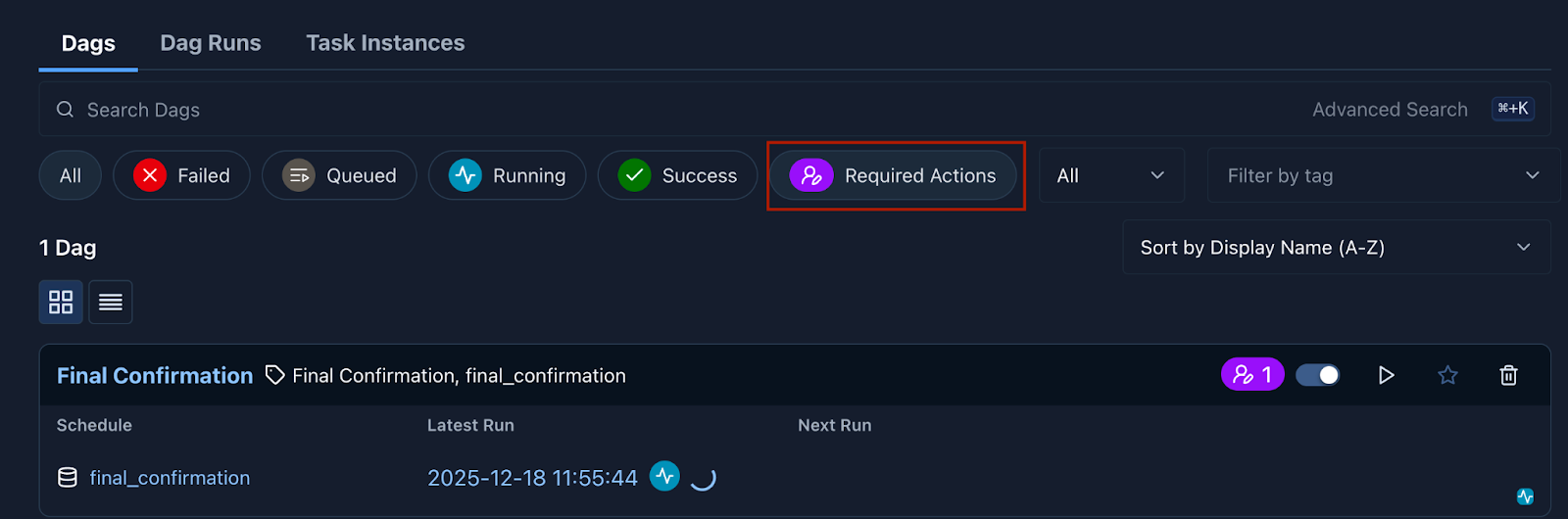
Figure 4: Notification of a Required Action in Airflow’s WebUI.
The required YAML file follows similar conventions to the typical container-config.yaml files used when submitting containers to MedPerf, but chains multiple containers in a single step. The basic structure is as follows, showing a single step. The mounts field in particular follows the exact same structure as the container-config.yaml files used in MedPerf for submitting Model and Metrics containers.
steps:
- id: unique_id_for_step
type: container
image: container_image_name:tag
command: command to run in container
mounts:
input_volumes:
data_path:
mount_path: /path/to/input/data/directory
type: directory
labels_path:
mount_path: /path/to/input/labels/directory
type: directory
parameters_file:
mount_path: /path/to/parameters.yaml
type: file
output_volumes:
output_path:
mount_path: /path/to/output/data/directory
type: directory
output_labels_path:
mount_path: /path/to/output/labels/directory
type: directory
next: id_for_next_step_or_null_if_last_step
What’s Next
The Apache Airflow integration is now available in MedPerf and is already running in production in the FL-PoST study. We invite the broader medical AI community to explore the implementation, test it in your own federated workflows and contribute back to the project.
– Explore the full implementation in the MedPerf GitHub repository
– Step-by-step YAML examples for building your own pipelines are available here.
– Interested in running a federated clinical study with MedPerf? Join our community.
The Medical Working Group continues to develop MedPerf as a community resource for reproducible, trustworthy AI evaluation in healthcare. Stay connected with MLCommons to follow future developments.
About MedPerf MedPerf is an open-source platform developed by the MLCommons Medical Working Group for benchmarking and evaluating AI/ML models in healthcare. Designed for federated clinical studies, MedPerf enables secure, reproducible AI evaluation across multiple sites without requiring raw patient data to leave local systems.
About MLCommons MLCommons is a non-profit engineering consortium dedicated to accelerating machine learning innovation for the benefit of humanity. Through open collaboration between industry, academia, and research institutions, MLCommons develops benchmarks, datasets, and best practices that advance AI safety, performance, and accessibility.
